Xenium human breast cancer brain - two samples
Melody Jin
2026-02-18
library(spatstat)
library(jazzPanda)
library(SpatialExperiment)
library(Seurat)
library(ggplot2)
library(data.table)
library(dplyr)
library(glmnet)
library(caret)
library(ComplexUpset)
library(corrplot)
library(gridExtra)
library(patchwork)
library(RColorBrewer)
library(limma)
library(edgeR)
library(speckle)
library(here)
source(here("code/utils.R"))Load sample1 and sample2 dataset
data_nm = "xenium_hbreast"
sp1 = get_xenium_data(rdata$xhb_r1,
mtx_name="cell_feature_matrix",
trans_name="transcripts.csv.gz",
cells_name="cells.csv.gz" )
sp1$trans_info = sp1$trans_info[sp1$trans_info$qv >=20 &
sp1$trans_info$cell_id != -1 &
!(sp1$trans_info$cell_id %in% sp1$zero_cells), ]
## sample2
sp2 = get_xenium_data(rdata$xhb_r2,
mtx_name="cell_feature_matrix",
trans_name="transcripts.csv.gz",
cells_name="cells.csv.gz" )
sp2$trans_info = sp2$trans_info[sp2$trans_info$qv >=20 &
sp2$trans_info$cell_id != -1 &
!(sp2$trans_info$cell_id %in% sp1$zero_cells), ]Single cell summary
These summary plots characterise per-cell detection counts and sparsity. We can calculate per-cell and per-gene summary statistics from the counts matrix:
Total detections per cell: sums all transcript counts in each cell.
Proportion of zeroes per cell: fraction of genes not detected in each cell.
Detected genes per cell: number of non-zero genes per cell.
Average expression per gene: mean expression across cells, ignoring zeros.
Each distribution is visualised with histograms and density curves to assess data quality and sparsity.
td_r1 <- colSums(sp1$cm)
pz_r1 <- colMeans(sp1$cm==0)
numgene_r1 <- colSums(sp1$cm!=0)
td_r2 <- colSums(sp2$cm)
pz_r2 <- colMeans(sp2$cm==0)
numgene_r2 <- colSums(sp2$cm!=0)
# Build the entire summary as one string
output <- paste0(
"\n================= Summary Statistics =================\n\n",
"--- Sample 1 ---\n",
make_sc_summary(td_r1, "Total detections per cell:"),
make_sc_summary(pz_r1, "Proportion of zeroes per cell:"),
make_sc_summary(numgene_r1, "Detected genes per cell:"),
"\n--- Sample 2 ---\n",
make_sc_summary(td_r2, "Total detections per cell:"),
make_sc_summary(pz_r2, "Proportion of zeroes per cell:"),
make_sc_summary(numgene_r2, "Detected genes per cell:"),
"\n========================================================\n"
)
cat(output)
================= Summary Statistics =================
--- Sample 1 ---
Total detections per cell:
Min: 1.0000 | 1Q: 97.0000 | Median: 166.0000 | Mean: 192.9693 | 3Q: 263.0000 | Max: 1359.0000
Proportion of zeroes per cell:
Min: 0.3802 | 1Q: 0.7476 | Median: 0.7955 | Mean: 0.7963 | 3Q: 0.8466 | Max: 0.9968
Detected genes per cell:
Min: 1.0000 | 1Q: 48.0000 | Median: 64.0000 | Mean: 63.7426 | 3Q: 79.0000 | Max: 194.0000
--- Sample 2 ---
Total detections per cell:
Min: 1.0000 | 1Q: 38.0000 | Median: 63.0000 | Mean: 89.7103 | 3Q: 101.0000 | Max: 1487.0000
Proportion of zeroes per cell:
Min: 0.4826 | 1Q: 0.8160 | Median: 0.8646 | Mean: 0.8583 | 3Q: 0.9062 | Max: 0.9965
Detected genes per cell:
Min: 1.0000 | 1Q: 27.0000 | Median: 39.0000 | Mean: 40.8172 | 3Q: 53.0000 | Max: 149.0000
========================================================td_df = as.data.frame(rbind(cbind(as.vector(td_r1),rep("sample1", length(td_r1))),
cbind(as.vector(td_r2),rep("sample2", length(td_r2)))))
colnames(td_df) = c("td","sample")
td_df$td= as.numeric(td_df$td)
p1<-ggplot(data = td_df, aes(x = td, color=sample)) +
geom_histogram(aes(y = after_stat(density)),
binwidth = 50, fill = "gray", color = "black") +
geom_density(color = "steelblue", size = 2) +
facet_wrap(~sample)+
labs(title = "Distribution of total detections per cell",
x = " ", y = "Density") +
#xlim(0, 1000) +
theme_minimal() +
theme(plot.title = element_text(hjust = 0.5),
strip.text = element_text(size=12))
pz = as.data.frame(rbind(cbind(as.vector(pz_r1),rep("sample1", length(pz_r1))),
cbind(as.vector(pz_r2),rep("sample2", length(pz_r2)))))
colnames(pz) = c("prop_ze","sample")
pz$prop_ze= as.numeric(pz$prop_ze)
p2<-ggplot(data = pz, aes(x = prop_ze, color=sample)) +
geom_histogram(aes(y = after_stat(density)),
binwidth = 0.05, fill = "gray", color = "black") +
geom_density(color = "steelblue", size = 2) +
facet_wrap(~sample)+
labs(title = "Distribution of proportion of zeroes per cell",
x = " ", y = "Density") +
#xlim(0, 1) +
theme_minimal() +
theme(plot.title = element_text(hjust = 0.5),
strip.text = element_text(size=12))
numgens = as.data.frame(rbind(cbind(as.vector(numgene_r1),rep("sample1", length(numgene_r1))),
cbind(as.vector(numgene_r2),rep("sample2", length(numgene_r2)))))
colnames(numgens) = c("numgen","sample")
numgens$numgen= as.numeric(numgens$numgen)
p3<-ggplot(data = numgens, aes(x = numgen, color=sample)) +
geom_histogram(aes(y = after_stat(density)),
binwidth = 10, fill = "gray", color = "black") +
geom_density(color = "steelblue", size = 2) +
facet_wrap(~sample)+
labs(title = "Distribution of detected genes per cell",
x = " ", y = "Density") +
theme_minimal() +
theme(plot.title = element_text(hjust = 0.5),
strip.text = element_text(size=12))cm_new1=sp1$cm
cm_new1[cm_new1==0] = NA
cm_new1 = as.data.frame(cm_new1)
cm_new1$avg2 = rowMeans(cm_new1,na.rm = TRUE)
summary(cm_new1$avg2) Min. 1st Qu. Median Mean 3rd Qu. Max.
1.02 1.34 1.71 2.13 2.38 13.50 cm_new2=sp2$cm
cm_new2[cm_new2==0] = NA
cm_new2 = as.data.frame(cm_new2)
cm_new2$avg2 = rowMeans(cm_new2,na.rm = TRUE)
summary(cm_new2$avg2) Min. 1st Qu. Median Mean 3rd Qu. Max.
1.01 1.24 1.45 1.82 1.90 13.43 avg_exp = as.data.frame(cbind("avg"=c(cm_new1$avg2, cm_new2$avg2),
"sample"=c(rep("sample1", nrow(sp1$cm)),
rep("sample2", nrow(sp1$cm)))))
avg_exp$avg=as.numeric(avg_exp$avg)
avg_exp$sample = factor(avg_exp$sample,
levels = c("sample1", "sample2"))
p4<-ggplot(data = avg_exp, aes(x = avg, color=sample)) +
geom_histogram(aes(y = after_stat(density)),
binwidth = 1, fill = "gray", color = "black") +
geom_density(color = "orange", linewidth = 1) +
facet_wrap(~sample)+
labs(title = "Distribution of average gene expression per cell",
x = " ", y = "Density") +
#xlim(0, 1) +
theme_minimal() +
theme(plot.title = element_text(hjust = 0.5),
strip.text = element_text(size=12))
layout_design <- (p1|p2)/(p3|p4)
layout_design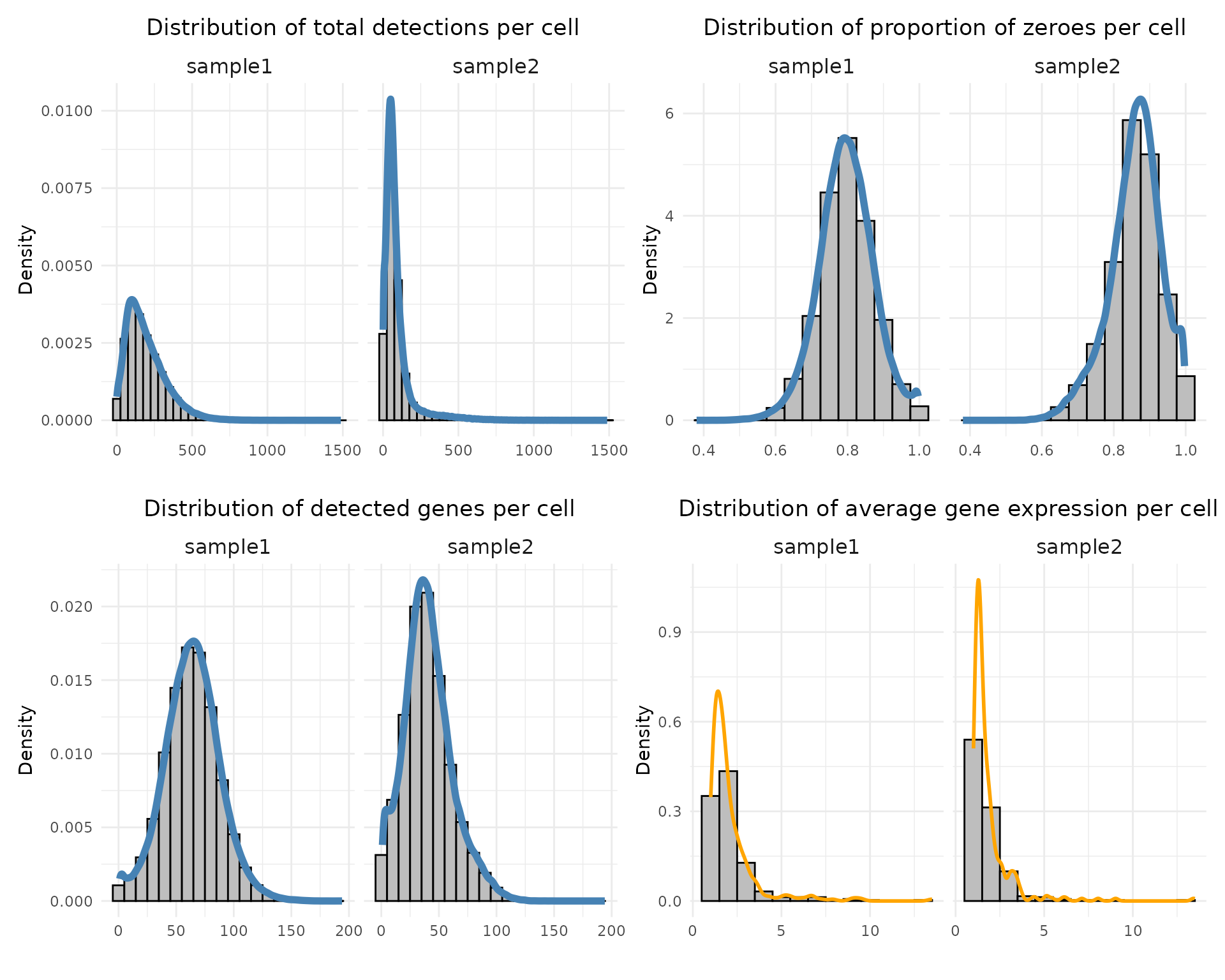
Both samples show similar overall distribution shapes across metrics. Sample 1 has higher transcript detection, with a median of 166 total counts and 64 detected genes per cell, while sample 2 shows lower detection levels (median 63 counts and 39 genes). The proportion of zeroes is high in both datasets (around 0.80 for sample 1 and 0.86 for sample 2). Average gene expression per cell is strongly right-skewed, indicating that most genes are expressed at low levels (median 1.71 in sample 1 and 1.45 in sample 2).
Quality assessment of negative controls
Negative control probes
Negative control probes are non-targeting probes designed not to hybridize to any transcript. There are 28 negative control probe targets in sample 1 and 20 in sample 2. In sample 1, the detection counts range from 110 to 12,975, with a median of 382, while in sample 2 they range from 15 to 86, with a median of 45. Notably, the probe antisense_PROKR2 shows a particularly high detection count (12,975) exclusively in sample 1.
probe_coords <- dplyr::bind_rows(
sp1$trans_info %>%
filter(feature_name %in% sp1$probe) %>%
select(feature_name, x, y) %>%
mutate(sample = "sample1"),
sp2$trans_info %>%
filter(feature_name %in% sp2$probe) %>%
select(feature_name, x, y) %>%
mutate(sample = "sample2")
)
ordered_feature = probe_coords %>% group_by(feature_name) %>% count() %>% arrange(desc(n))%>% pull(feature_name)
probe_tb = as.data.frame(probe_coords %>% group_by(sample, feature_name) %>% count())
colnames(probe_tb) = c("sample","feature_name","value_count")
probe_tb$feature_name = factor(probe_tb$feature_name, levels= ordered_feature)
probe_tb = probe_tb[order(probe_tb$feature_name), ]
p1<- ggplot(probe_tb, aes(x = feature_name, y = value_count)) +
geom_bar(stat = "identity", position = position_dodge()) +
facet_wrap(~sample, ncol=2, scales = "free")+
labs(title = " ", x = " ", y = "Number of total detections") +
theme_bw()+
theme(legend.position = "none",
axis.text.x= element_text(size=8, angle=45,vjust=1,hjust = 1),
aspect.ratio = 5/7,
axis.text.y=element_text(size=8),
axis.ticks=element_blank(),
axis.title.x=element_blank(),
strip.background = element_rect(fill="white", colour="black"),
panel.background=element_blank(),
panel.grid.minor=element_blank(),
plot.background=element_blank())
p1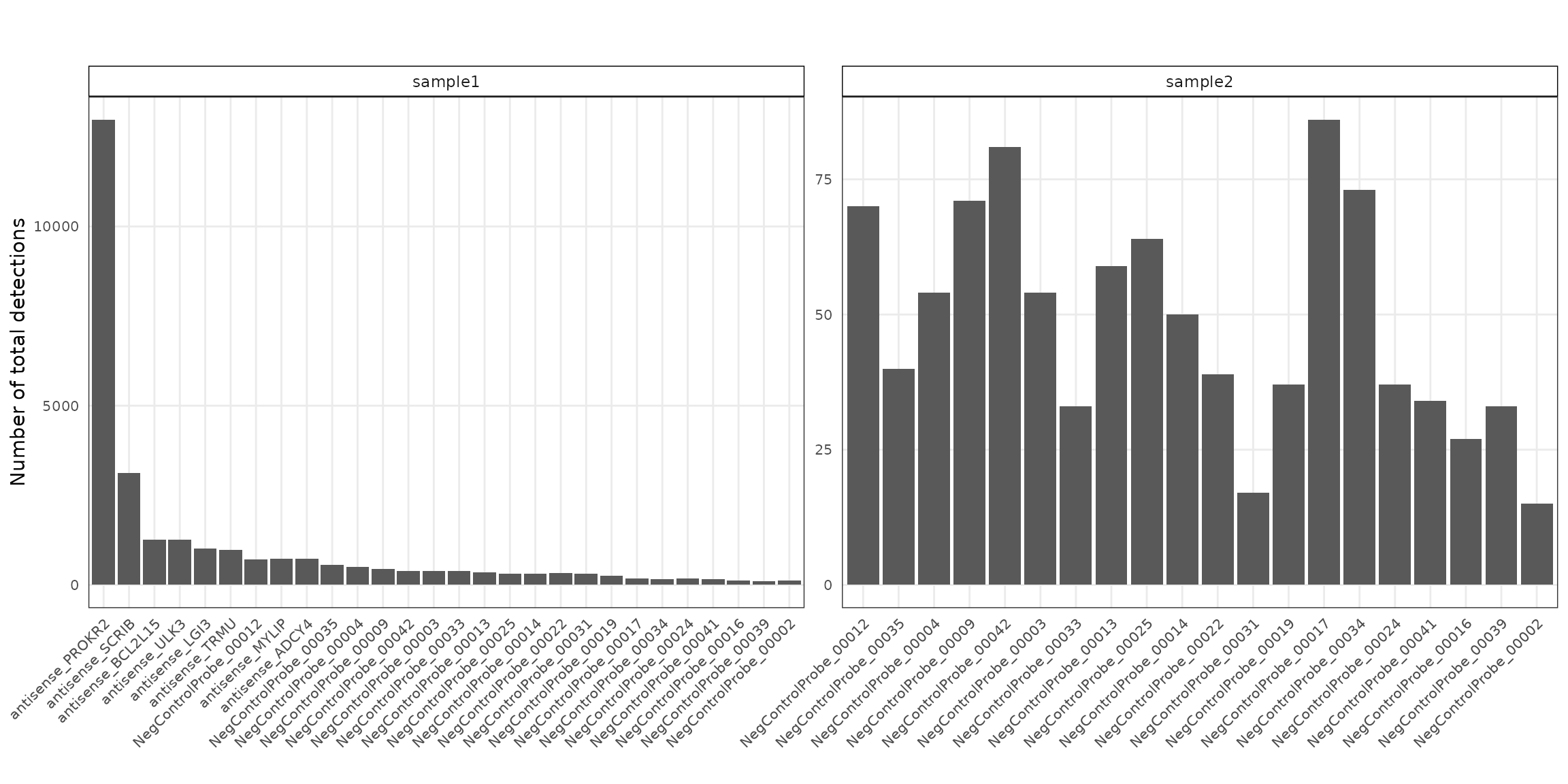
p1<-ggplot(probe_coords, aes(x = x, y = y)) +
geom_hex(bins = auto_hex_bin(min(table(probe_coords$sample)),
target_points_per_bin = 1)) +
theme_bw() +
scale_fill_gradient(low = "white", high = "black") +
guides(fill = guide_colorbar(barheight = unit(0.06, "npc"),
barwidth = unit(0.4, "npc"),
)) +
scale_x_reverse()+
facet_wrap(~sample, ncol=2) +
defined_theme +
theme(legend.title = element_text(size = 10,
hjust = 1, vjust=0.8),
legend.position = "bottom",
aspect.ratio = 7/10,
legend.text = element_text(size = 9),
strip.text = element_text(size = 12),
strip.background = element_blank())
p1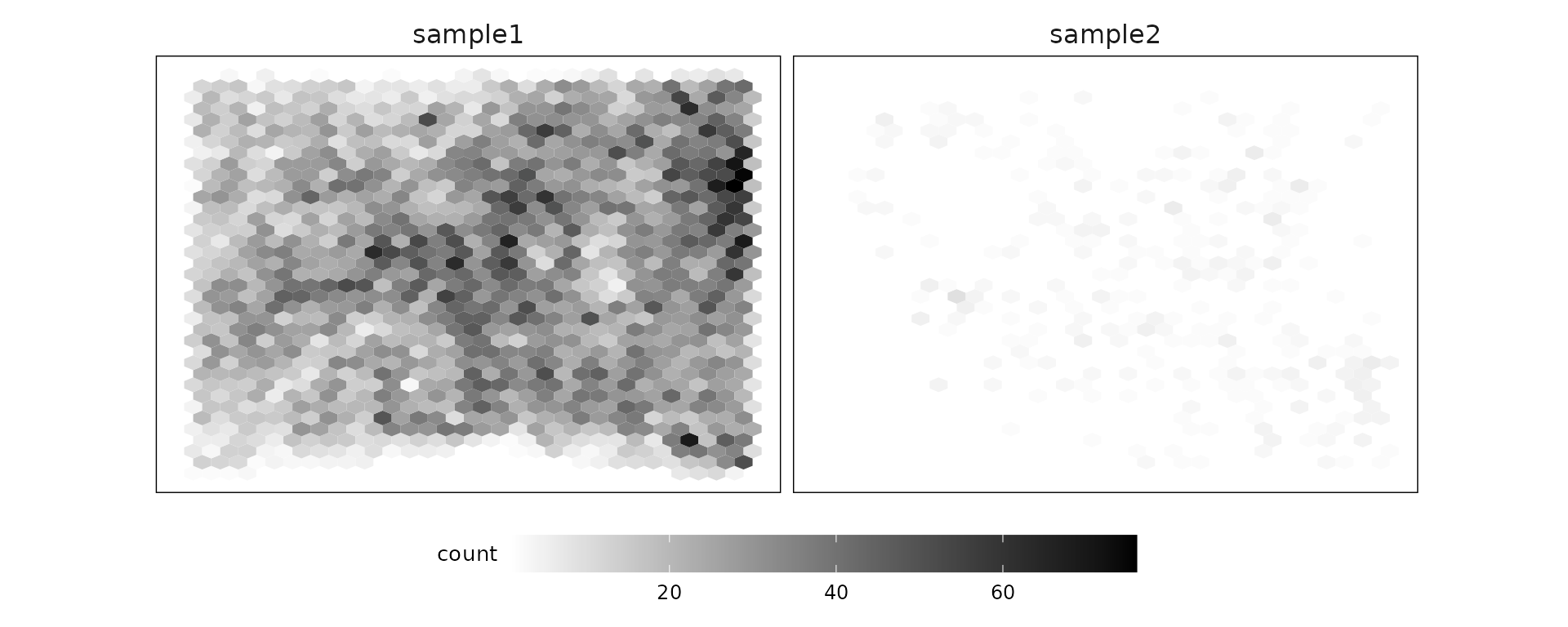
Negative control codewords
Negative control codewords are unused barcode sequences included to detect random or erroneous decoding events, providing a measure of decoding noise. In this dataset, 41 negative codeword targets were used, with median detection counts of about 46 in sample 1 and 6 in sample 2. There are no particular localised spatial patterns for these codeword targets.
codeword_coords <- dplyr::bind_rows(
sp1$trans_info %>%
filter(feature_name %in% sp1$codeword) %>%
select(feature_name, x, y) %>%
mutate(sample = "sample1"),
sp2$trans_info %>%
filter(feature_name %in% sp2$codeword) %>%
select(feature_name, x, y) %>%
mutate(sample = "sample2")
)
ordered_feature = codeword_coords %>%
group_by(feature_name) %>%
count() %>%
arrange(desc(n)) %>% pull(feature_name)
codeword_tb = as.data.frame(codeword_coords %>%
group_by(sample, feature_name) %>%
count())
colnames(codeword_tb) = c("sample","feature_name","value_count")
codeword_tb$feature_name = factor(codeword_tb$feature_name, levels= ordered_feature)
codeword_tb = codeword_tb[order(codeword_tb$feature_name), ]
p2<- ggplot(codeword_tb, aes(x = feature_name, y = value_count)) +
geom_bar(stat = "identity", position = position_dodge()) +
facet_wrap(~sample, ncol=2, scales = "free")+
labs(title = " ", x = " ", y = "Number of total detections") +
theme_bw()+
theme(legend.position = "none",
axis.text.x= element_text(size=8, angle=45,vjust=1,hjust = 1),
aspect.ratio = 5/7,
axis.text.y=element_text(size=8),
axis.ticks=element_blank(),
axis.title.x=element_blank(),
strip.background = element_rect(fill="white", colour ="black"),
panel.background=element_blank(),
panel.grid.minor=element_blank(),
plot.background=element_blank())
p2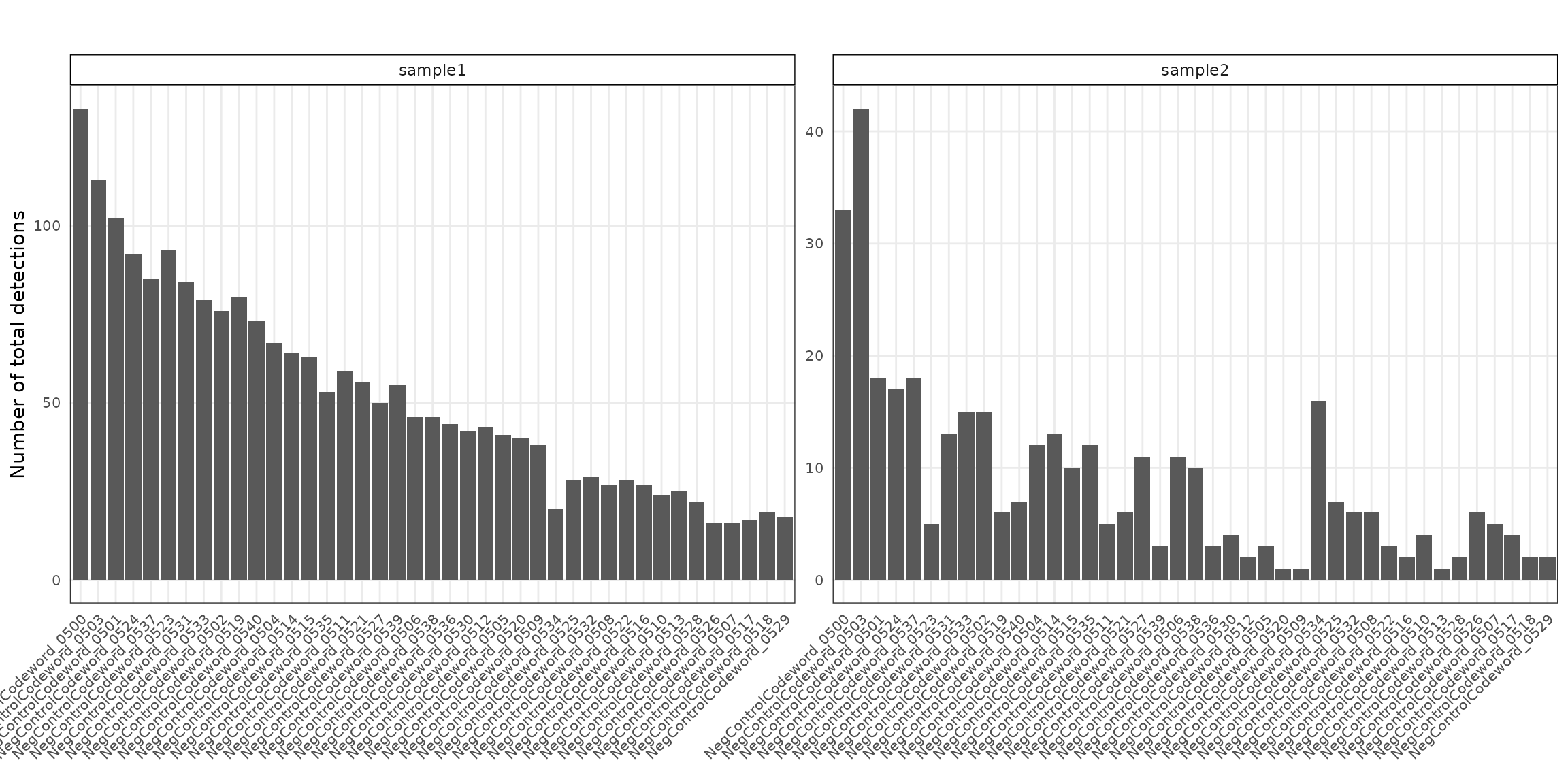
p2<- ggplot(codeword_coords, aes(x = x, y = y)) +
geom_hex(bins = auto_hex_bin(min(table(probe_coords$sample)),
target_points_per_bin = 1)) +
theme_bw() +
scale_fill_gradient(low = "white", high = "black") +
guides(fill = guide_colorbar(barheight = unit(0.06, "npc"),
barwidth = unit(0.4, "npc"),
)) +
scale_x_reverse()+
facet_wrap(~sample, ncol=2) +
defined_theme +
theme(legend.title = element_text(size = 10, hjust = 1, vjust=0.8),
legend.position = "bottom",
aspect.ratio = 7/10,
legend.text = element_text(size = 9),
strip.text = element_text(size = 12),
strip.background = element_blank())
p2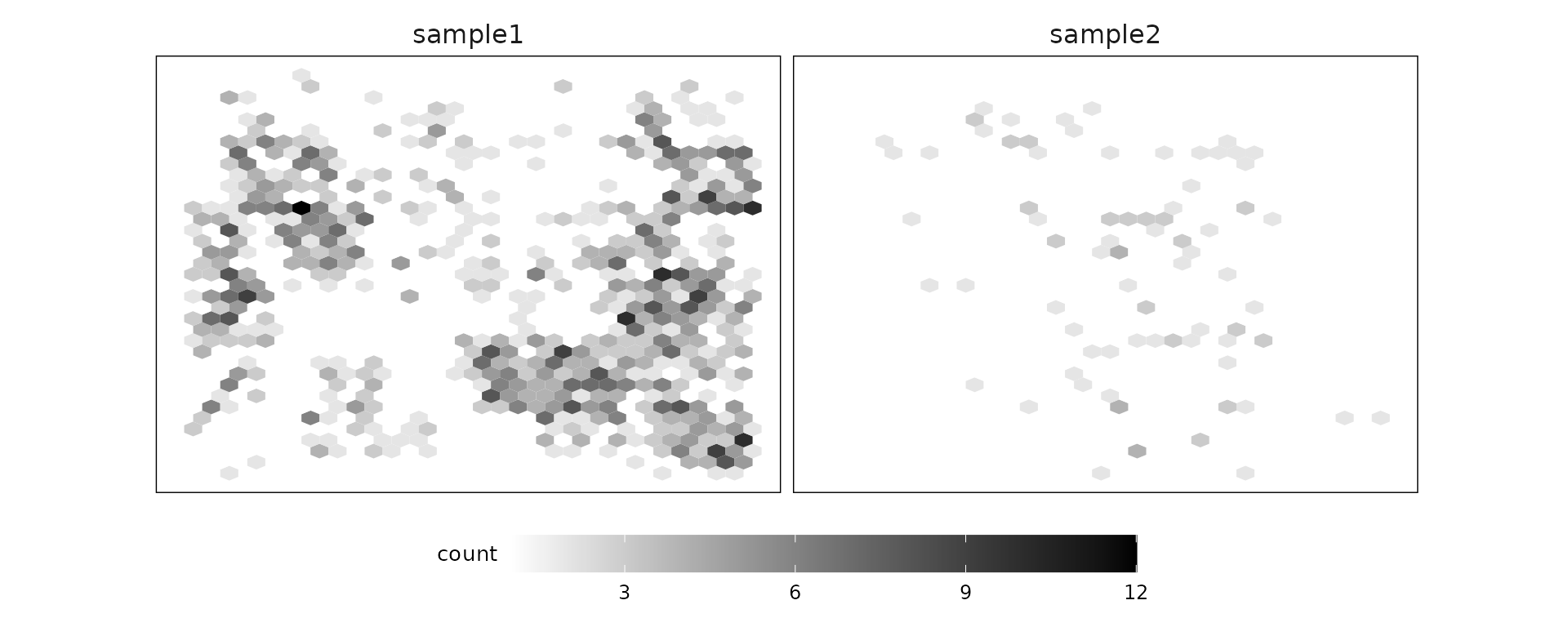
Clustering
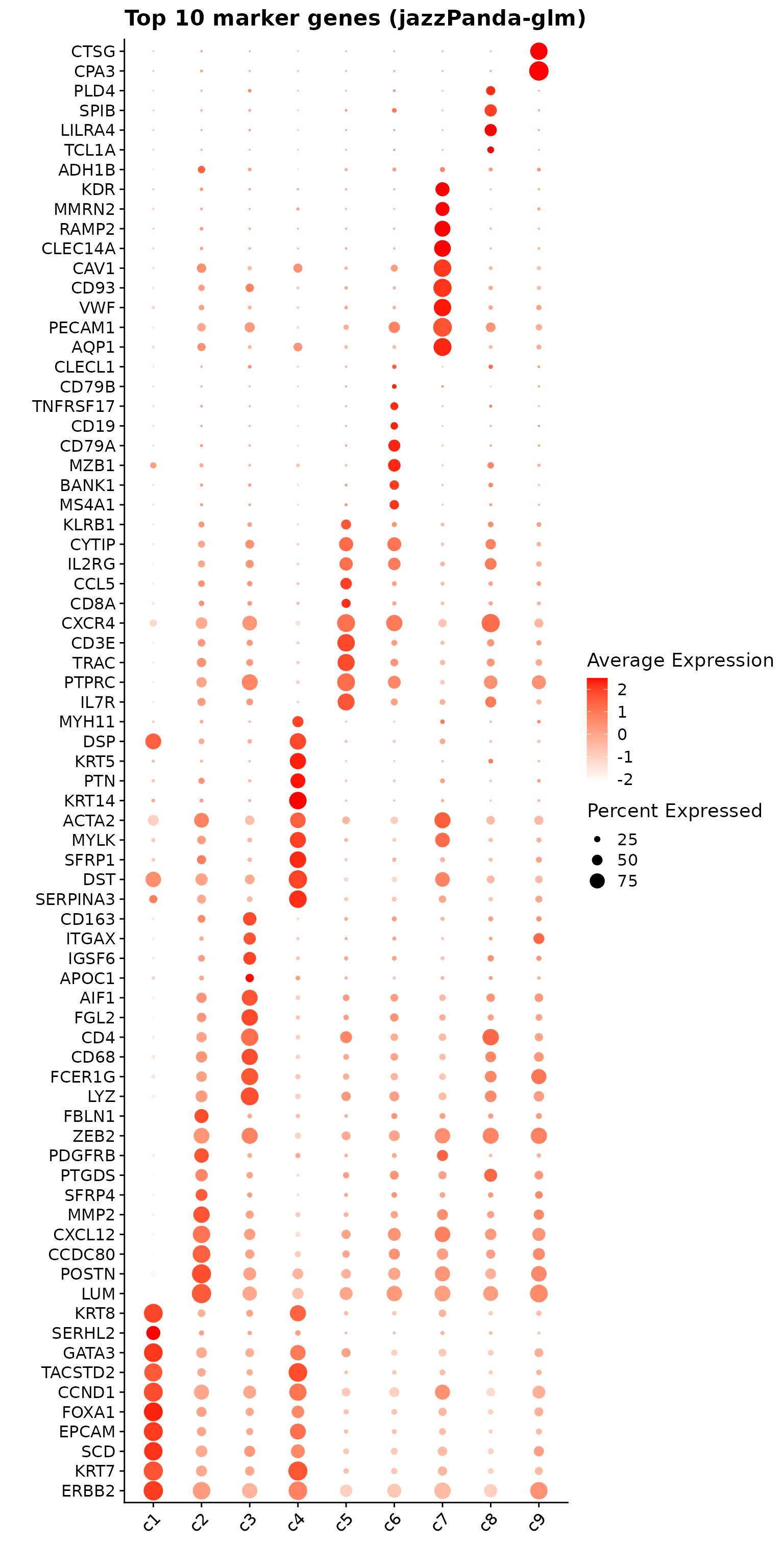
We process the two samples separately, and align them based on the shared cell types. For sample 1, we used the provided cell type labels and refined them by collapsing subtypes. For sample 2, we perform Louvain clustering with resolution 0.5, and annotate the resulting clusters with known marker genes. This process produced 14 cell types for sample 2, which were then grouped into broader categories for alignment with sample 1. Finally, cells from nine shared cell types in each sample were joined, and a cluster label (c1-c9) was created for representing each of the cell type.
project_root="/path/to/jazzPanda_paper/dir"
graphclust_sp1=read.csv(file.path(project_root, "data", "Xenium_rep1_supervised_celltype.csv"))
graphclust_sp1$anno =as.character(graphclust_sp1$Cluster)
target_clusters = c("Tumor", "Stromal","Macrophages","Myoepithelial",
"T_Cells", "B_Cells","Endothelial", "Dendritic", "Mast_Cells")
t_cells = c("CD4+_T_Cells","CD8+_T_Cells","T_Cell_&_Tumor_Hybrid",
"Stromal_&_T_Cell_Hybrid")
dc_cells = c("LAMP3+_DCs","IRF7+_DCs")
macro_cells = c("Macrophages_1","Macrophages_2")
myo_cells = c("Myoepi_KRT15+", "Myoepi_ACTA2+")
tumor_cells = c("Invasive_Tumor", "Prolif_Invasive_Tumor")
graphclust_sp1[graphclust_sp1$Cluster %in% t_cells, "anno"] = "T_Cells"
graphclust_sp1[graphclust_sp1$Cluster %in% macro_cells,"anno"] = "Macrophages"
graphclust_sp1[graphclust_sp1$Cluster %in% c("DCIS_1", "DCIS_2"),"anno"] = "DCIS"
graphclust_sp1[graphclust_sp1$Cluster %in% dc_cells,"anno"] = "Dendritic"
graphclust_sp1[graphclust_sp1$Cluster %in% myo_cells,"anno"] = "Myoepithelial"
graphclust_sp1[graphclust_sp1$Cluster %in% tumor_cells,"anno"] = "Tumor"
graphclust_sp1$anno=factor(graphclust_sp1$anno,
levels=c("Tumor", "DCIS", "Stromal",
"Macrophages","Myoepithelial",
"T_Cells", "B_Cells",
"Endothelial",
"Dendritic", "Mast_Cells",
"Perivascular-Like","Unlabeled"))
sp1_clusters = as.data.frame(cbind(as.character(graphclust_sp1$anno),
paste("c",as.numeric(factor(graphclust_sp1$anno)),
sep="")))
row.names(sp1_clusters) = graphclust_sp1$Barcode
colnames(sp1_clusters) = c("anno","cluster")
cells= sp1$cell_info
row.names(cells) =cells$cell_id
rp_names = row.names(sp1_clusters)
sp1_clusters[rp_names,"x"] = cells[match(rp_names,row.names(cells)),
"x"]
sp1_clusters[rp_names,"y"] = cells[match(rp_names,row.names(cells)),
"y"]
sp1_clusters$anno=factor(sp1_clusters$anno,
levels=c("Tumor", "DCIS", "Stromal",
"Macrophages","Myoepithelial",
"T_Cells", "B_Cells",
"Endothelial",
"Dendritic", "Mast_Cells",
"Perivascular-Like","Unlabeled"))
sp1_clusters$cluster=factor(sp1_clusters$cluster,
levels=paste("c", 1:12, sep=""))
sp1_clusters$cells =paste(row.names(sp1_clusters),"_1",sep="")
sp1_clusters$sample="sample1"
selected_sp1 = sp1_clusters
selected_sp1$cluster_comb = as.character(selected_sp1$anno)
selected_sp1[selected_sp1$cluster_comb =="DCIS","cluster_comb"] = "Tumor"
selected_sp1 = selected_sp1[selected_sp1$cluster_comb %in% target_clusters,]
selected_sp1$cluster_comb = factor(selected_sp1$cluster_comb, levels=target_clusters)
selected_sp1$cluster =paste("c",as.numeric(factor(selected_sp1$cluster_comb)),
sep="")
selected_sp1 = selected_sp1[,c("cluster","x","y","cells","sample","cluster_comb")]
colnames(selected_sp1)[6]="anno"
sample2_anno = read.csv(file.path(project_root, "data", "Sample2_Xenium_cell_type_manual.csv"),
row.names = 1)
sp2_clusters = as.data.frame(cbind(row.names(sample2_anno),
as.character(sample2_anno$cell_type)))
colnames(sp2_clusters) = c("cells","anno")
row.names(sp2_clusters) = sp2_clusters$cells
cells= sp2$cell_info
row.names(cells) = cells$cell_id
rp_names = sp2_clusters$cells
sp2_clusters[rp_names,"x"] = cells[match(rp_names,row.names(cells)),
"x"]
sp2_clusters[rp_names,"y"] = cells[match(rp_names,row.names(cells)),
"y"]
sp2_clusters$sample="sample2"
sp2_clusters[sp2_clusters$anno %in% c("B", "Plasma"),"anno"]="B_Cells"
sp2_clusters[sp2_clusters$anno %in% c("Macrophage"),"anno"]="Macrophages"
sp2_clusters[sp2_clusters$anno %in% c("T","NK"),"anno"]="T_Cells"
sp2_clusters[sp2_clusters$anno %in% c("Fibroblast"),"anno"]="Stromal"
sp2_clusters[sp2_clusters$anno %in% c("Mast"),"anno"]="Mast_Cells"
sp2_clusters$anno = factor(sp2_clusters$anno,
levels=c("Tumor", "Stromal","Macrophages",
"Myoepithelial", "T_Cells", "B_Cells",
"Endothelial", "Dendritic", "Mast_Cells",
"ML","LP","Adipocyte"))
sp2_clusters$cells=paste(sp2_clusters$cells,"-sp2", sep="")
sp2_clusters$anno = as.character(sp2_clusters$anno)
selected_sp2 = sp2_clusters[sp2_clusters$anno %in% target_clusters,]
selected_sp2$anno = factor(selected_sp2$anno, levels=target_clusters)
selected_sp2$cluster =paste("c",as.numeric(factor(selected_sp2$anno)),
sep="")
selected_sp2 = selected_sp2[,c("cluster","x","y","cells","sample","anno")]
clusters = rbind(selected_sp1, selected_sp2)
table(clusters$sample, clusters$anno)
clusters$anno = factor(clusters$anno, target_clusters)
saveRDS(cluster_info, here(gdata,paste0(data_nm, "_clusters.Rds")))xhb_color<- c("#FC8D62","#66C2A5" ,"#8DA0CB","#E78AC3",
"#A6D854","skyblue","purple3","#E5C498","blue")
# load generated data (see code above for how they were created)
cluster_info = readRDS(here(gdata,paste0(data_nm, "_clusters.Rds")))
colnames(cluster_info)[6] = "anno_name"
cluster_names = paste0("c", 1:9)
cluster_info$cluster = factor(cluster_info$cluster,
levels=cluster_names)
ct_names =c("Tumor", "Stromal","Macrophages","Myoepithelial", "T_Cells",
"B_Cells","Endothelial", "Dendritic", "Mast_Cells")
cluster_info$anno_name = factor(cluster_info$anno_name,
levels=ct_names)
cluster_info = cluster_info[order(cluster_info$anno_name), ]
cluster_info$cells = paste0("_", cluster_info$cells)
anno_df = unique(cluster_info[c("cluster", "anno_name")])
anno_df$anno_name = factor(anno_df$anno_name, levels = ct_names)We can see that While both samples contain the same major cell types, their relative proportions differ markedly. Sample 1 is dominated by tumor and stromal cells, which together account for more than 65% of cells. In contrast, sample 2 shows a lower tumor and stromal fraction but a clear increase in immune and vascular components. Notably, T cells are more than twice as abundant in sample 2, accompanied by higher proportions of B cells, endothelial cells, dendritic cells, and mast cells.
sum_df = cluster_info[,c("anno_name", "cells","sample"),drop=FALSE] %>%
group_by(anno_name, sample) %>% dplyr::count()
ggplot(sum_df, aes(x = sample, y = n, fill=anno_name)) +
geom_bar( stat="identity",position="fill") +
scale_y_continuous(labels = scales::percent)+
labs(title = " ", x = " ", y = "proportion of cell types", fill="cell types") +
scale_fill_manual(values =xhb_color)+
theme(legend.position = "right",axis.text.x= element_text(size=12),
strip.text = element_text(size = rel(1)),
axis.line=element_blank(),
axis.text.y=element_blank(),axis.ticks=element_blank(),
axis.title.x=element_blank(),
axis.title.y=element_blank(),
panel.background=element_blank(),
panel.grid.major=element_blank(),
panel.grid.minor=element_blank(),
plot.background=element_blank())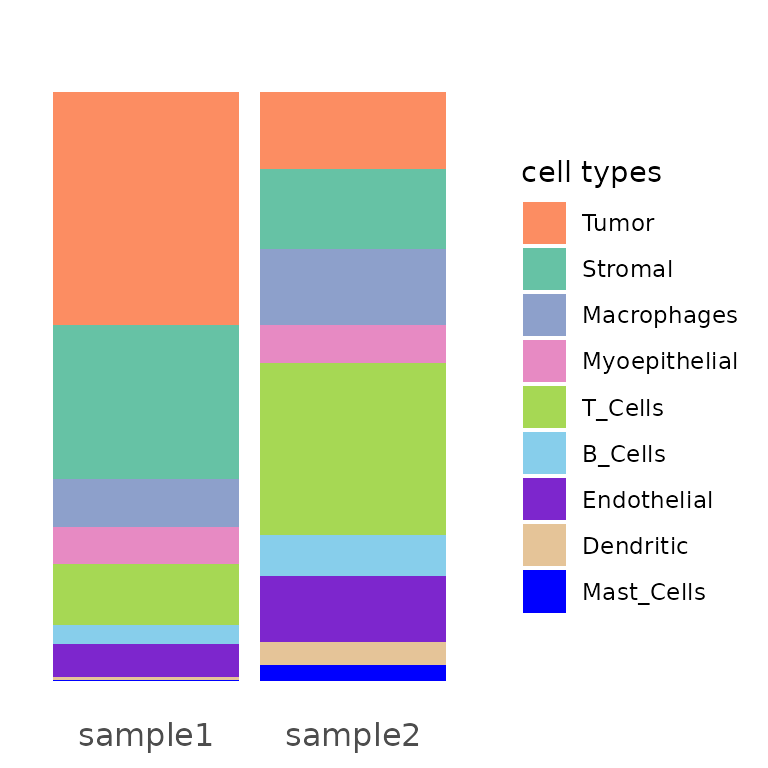
ggplot(data = cluster_info,
aes(x = x, y = y, color=cluster))+
geom_point(size=0.001)+
facet_wrap(~sample, nrow=1)+
theme_classic()+
scale_y_reverse()+
scale_color_manual(values =xhb_color)+
guides(color=guide_legend(title="cluster", ncol = 1,
override.aes=list(alpha=1, size=8)))+ defined_theme+
theme(strip.background = element_rect(color="white"),
legend.position = "right",legend.key.width = unit(1, "cm"),
aspect.ratio = 5/7,
legend.title = element_text(size = 12),
legend.text = element_text(size = 12),
legend.key.height = unit(1, "cm"),
strip.text = element_text(size = rel(2.2)))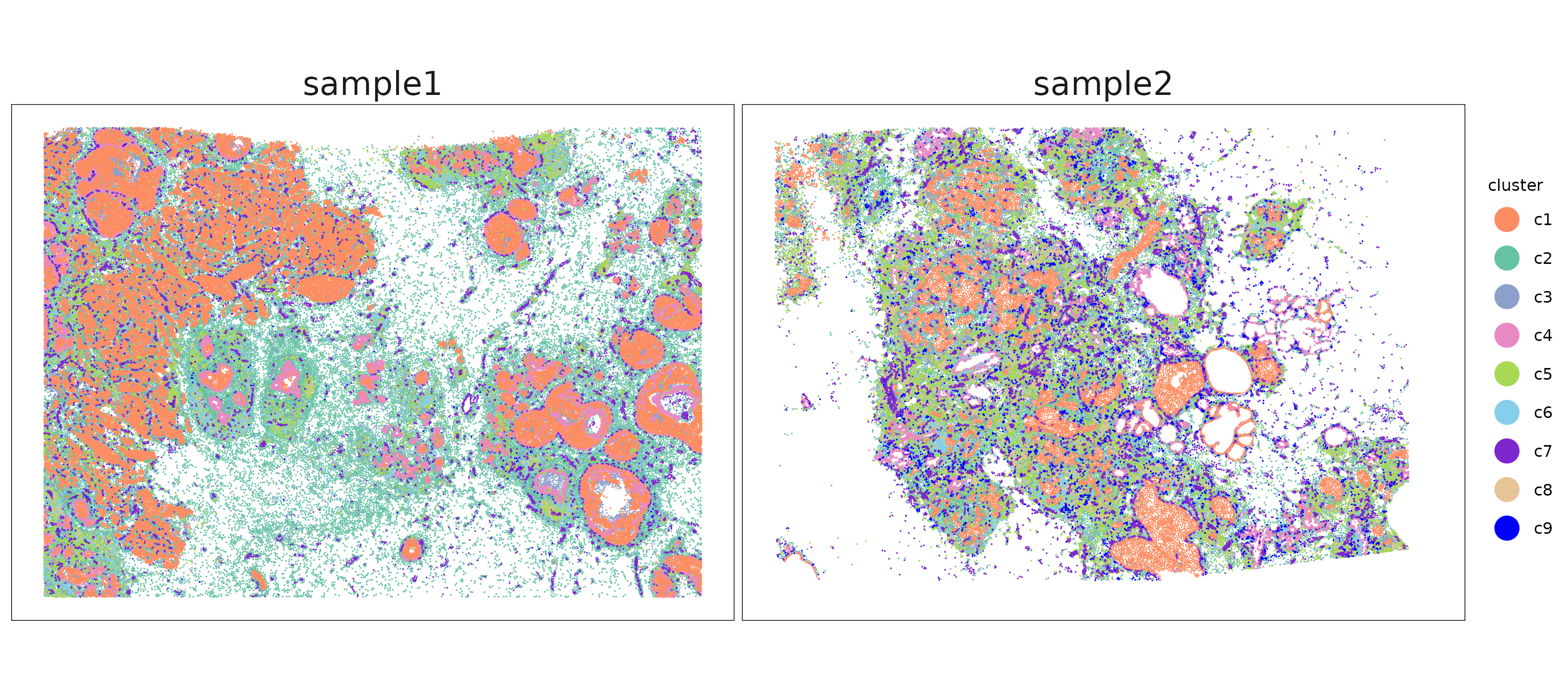
The clusters show distinct and largely non-overlapping spatial patterns. We can see that the corresponding clusters in sample 1 and sample 2 differ in their spatial arrangement, reflecting heterogeneity in tissue structures.
p1<- ggplot(data = cluster_info[cluster_info$sample=="sample1", ],
aes(x = x, y = y, color=cluster))+
geom_point(size=0.001)+
facet_wrap(~cluster, ncol=3)+
scale_y_reverse()+
scale_color_manual(values = xhb_color)+
theme_classic()+
guides(color=guide_legend(title="cluster",
override.aes=list(alpha=1, size=8)))+
theme_minimal()+
defined_theme+
labs(title = "Sample1")+
theme(aspect.ratio = 5/7,
plot.title = element_text(size = rel(1.6)),
strip.text = element_text(size = rel(1.2)))
p2<- ggplot(data = cluster_info[cluster_info$sample=="sample2", ],
aes(x = x, y = y, color=cluster))+
geom_point(size=0.001)+
facet_wrap(~cluster, ncol=3)+
scale_y_reverse()+
scale_color_manual(values = xhb_color)+
theme_classic()+
guides(color=guide_legend(title="cluster",
override.aes=list(alpha=1, size=8)))+
theme_minimal()+
defined_theme+
labs(title = "Sample2")+
theme(aspect.ratio = 5/7,
plot.title = element_text(size = rel(1.6)),
strip.text = element_text(size = rel(1.2)))
layout_design <- p1 / p2
layout_design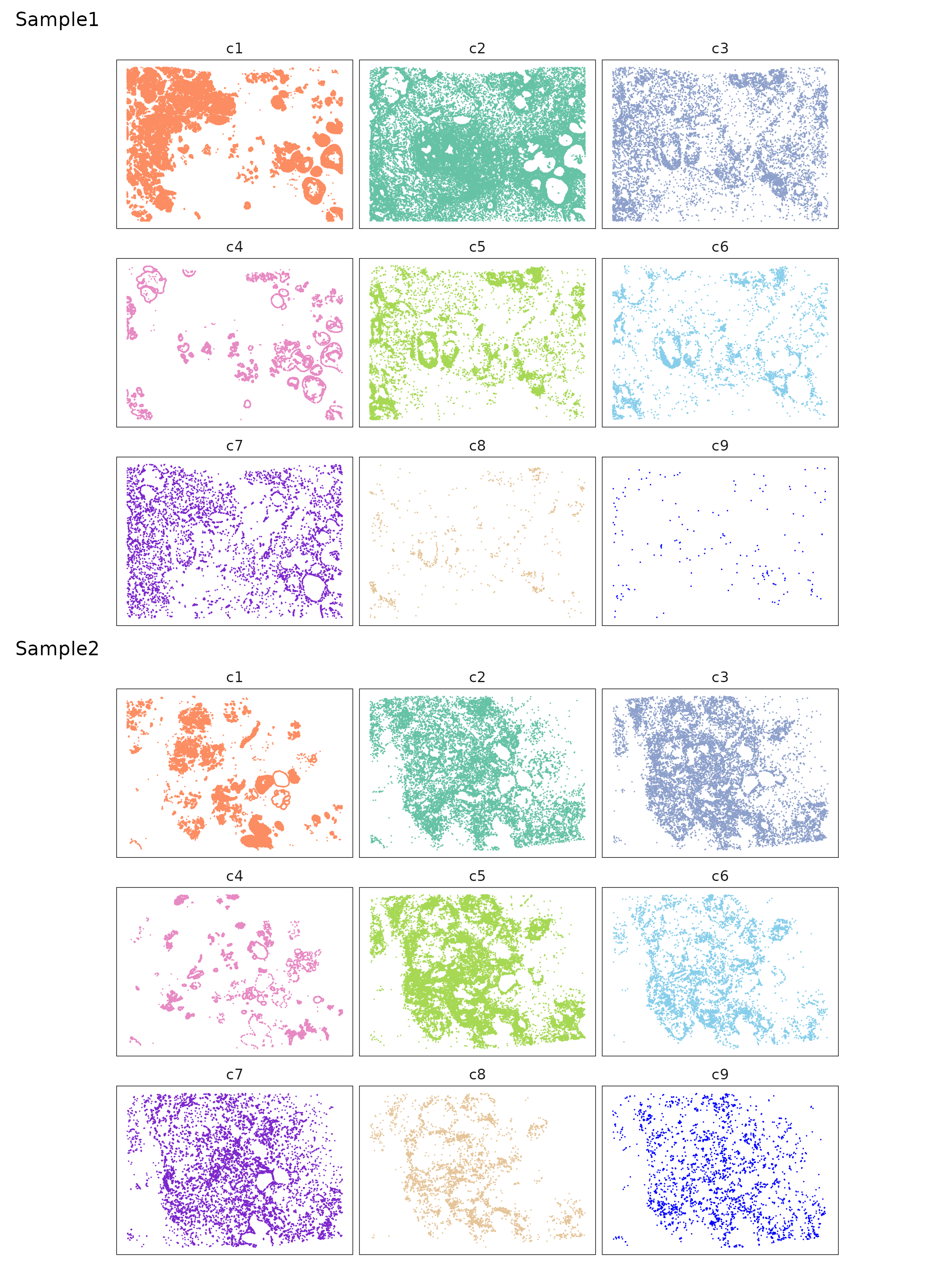
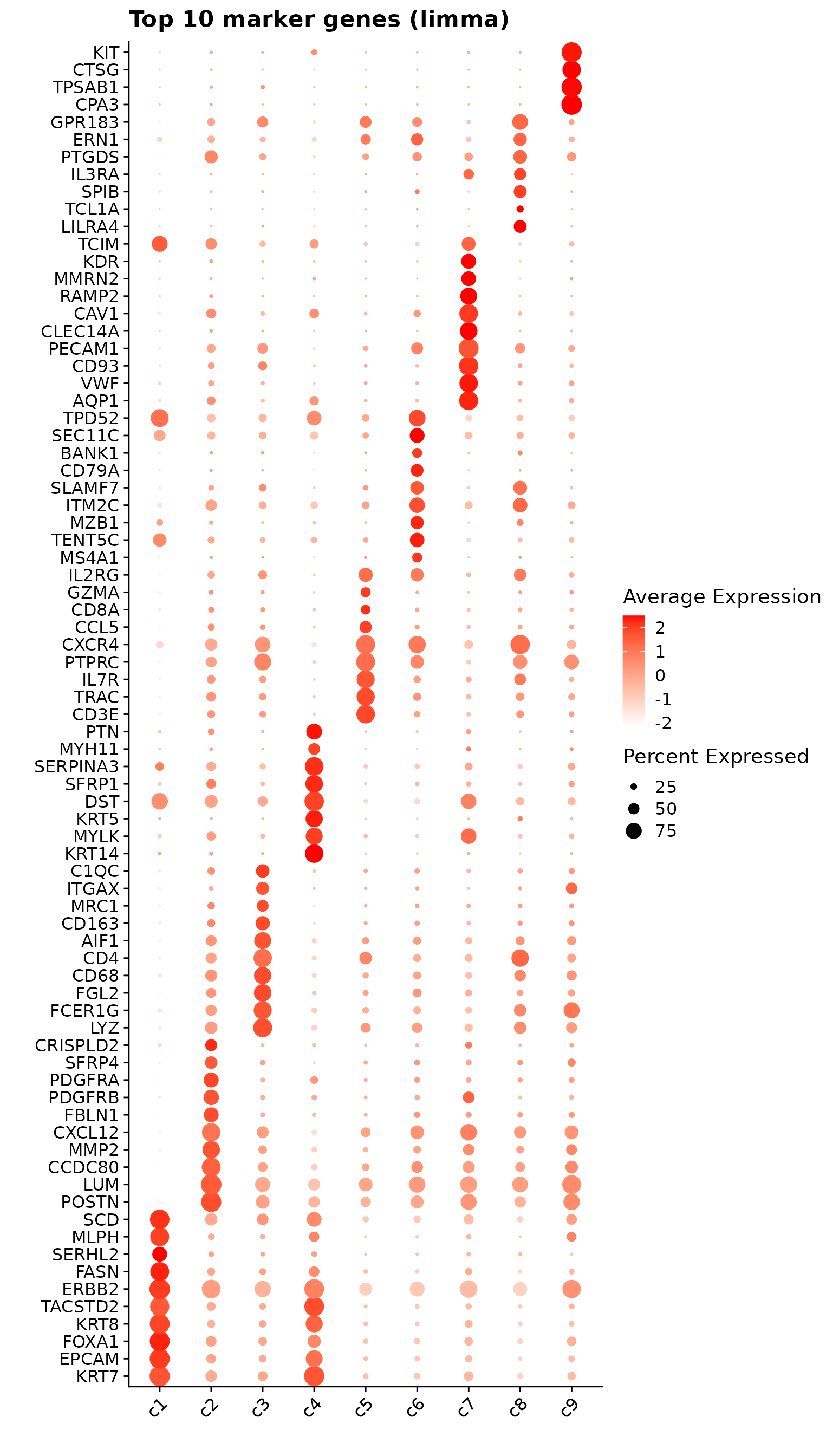
Comparison of marker gene detection methods
Cumulative rank plot
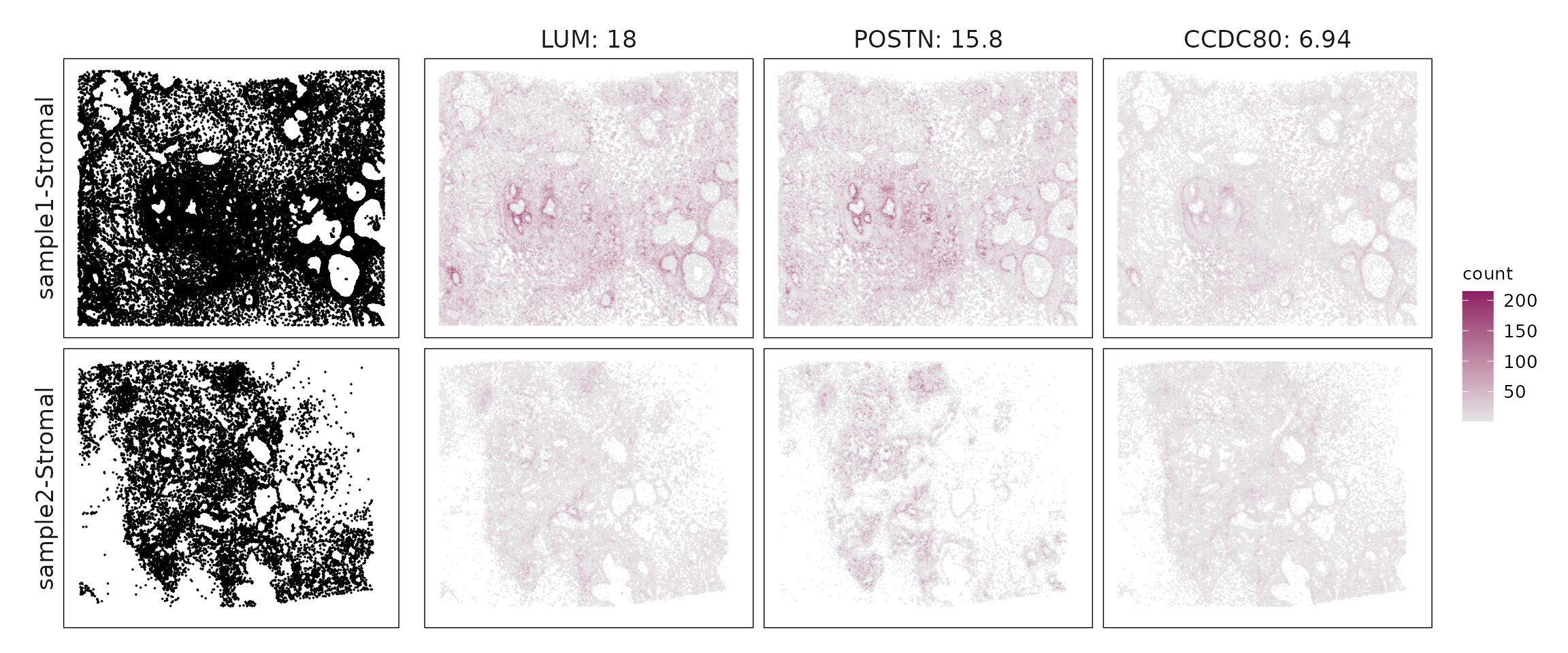
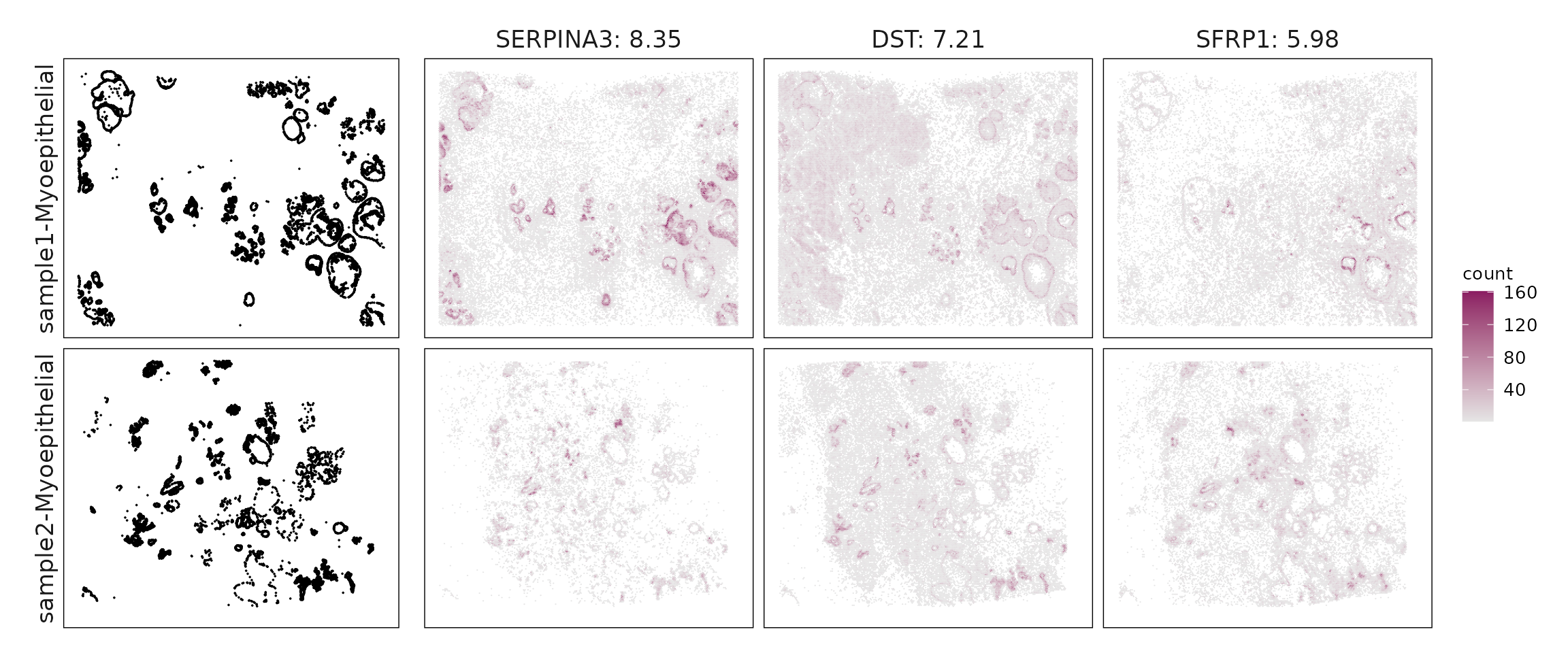
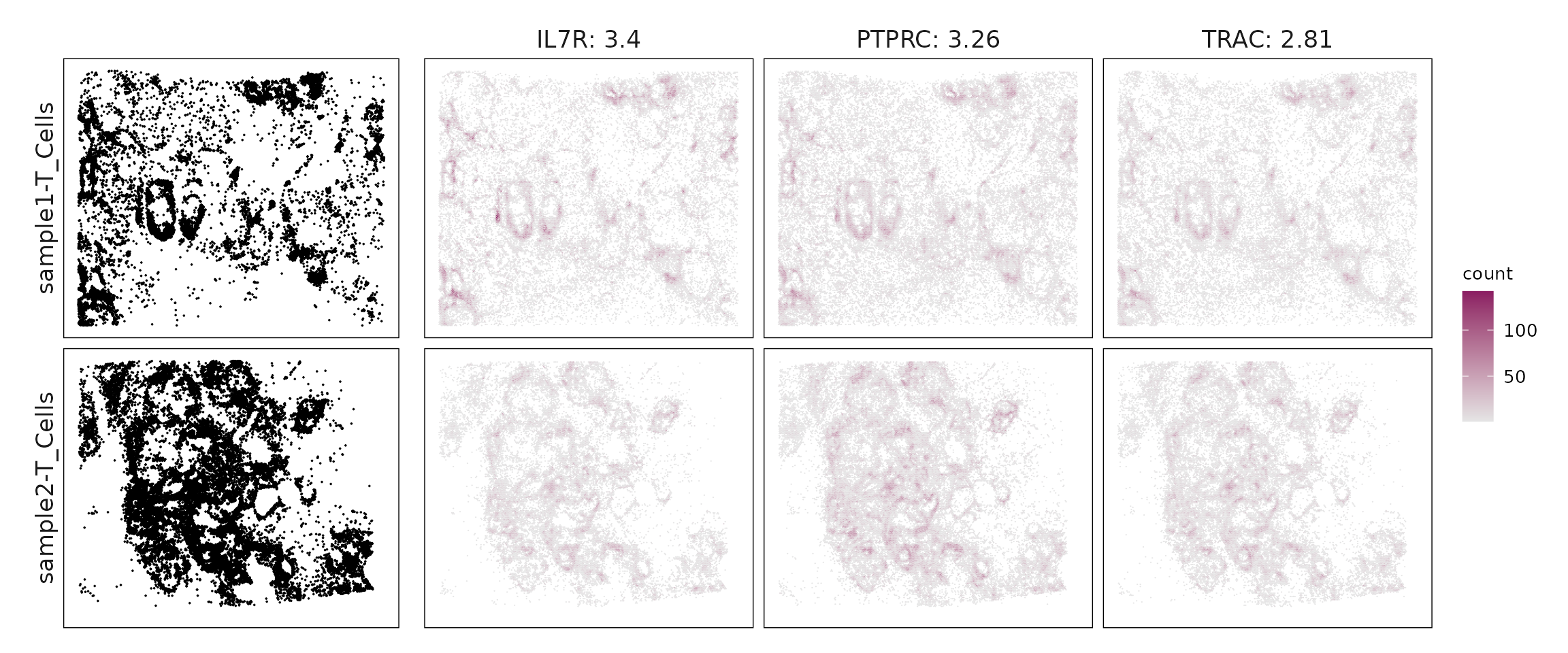
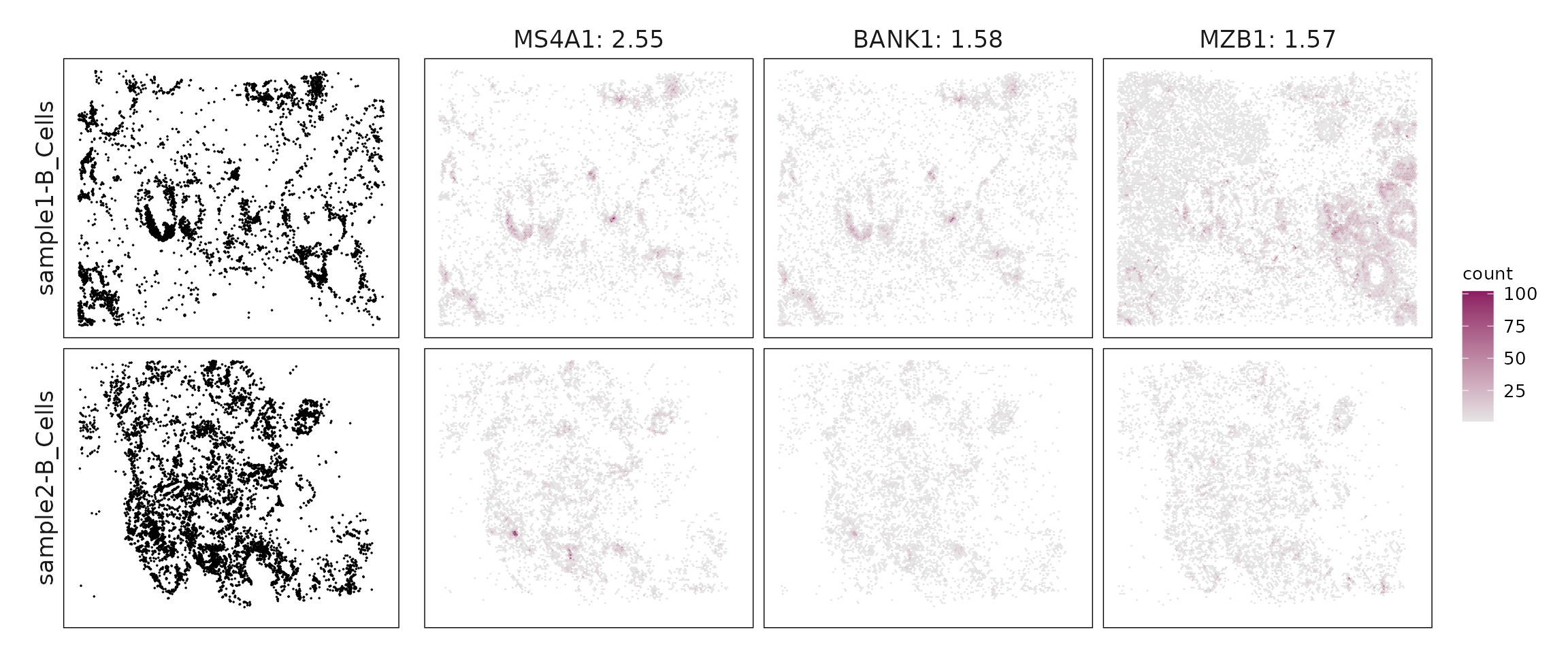
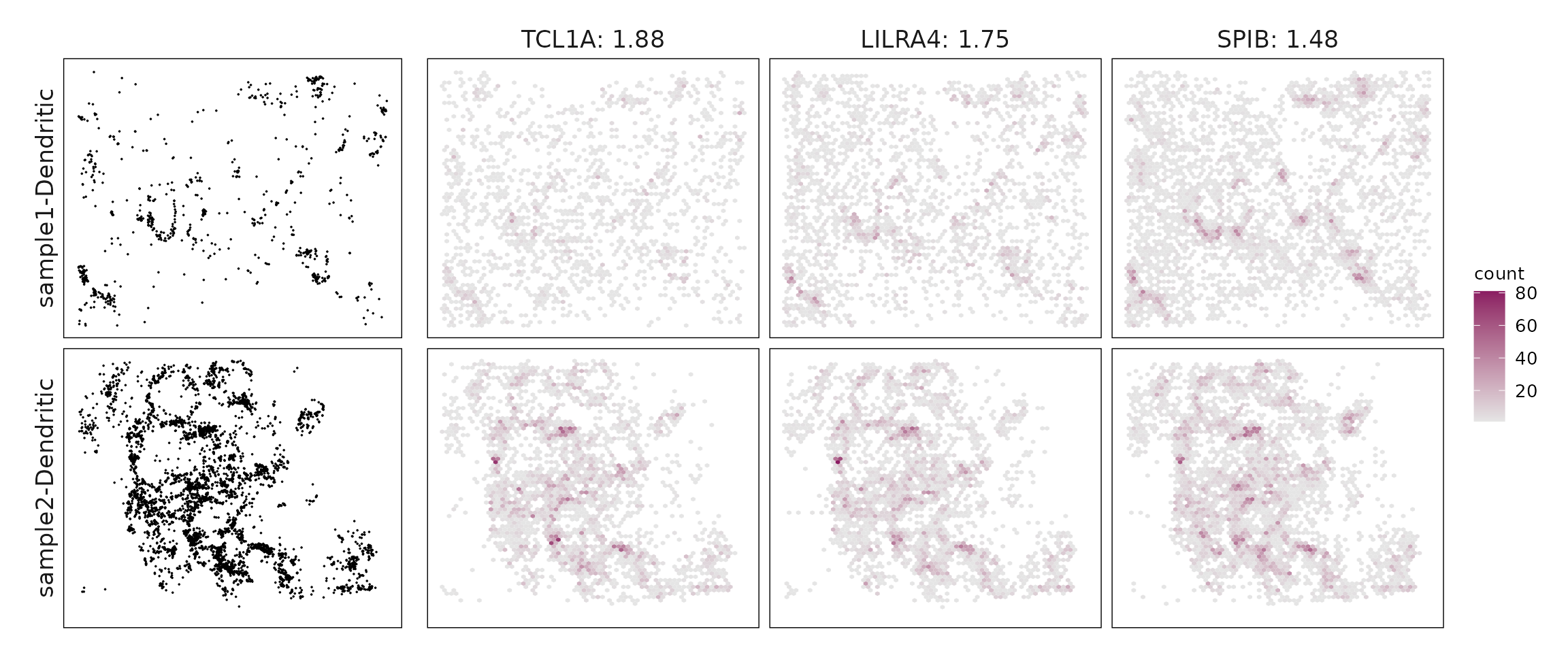
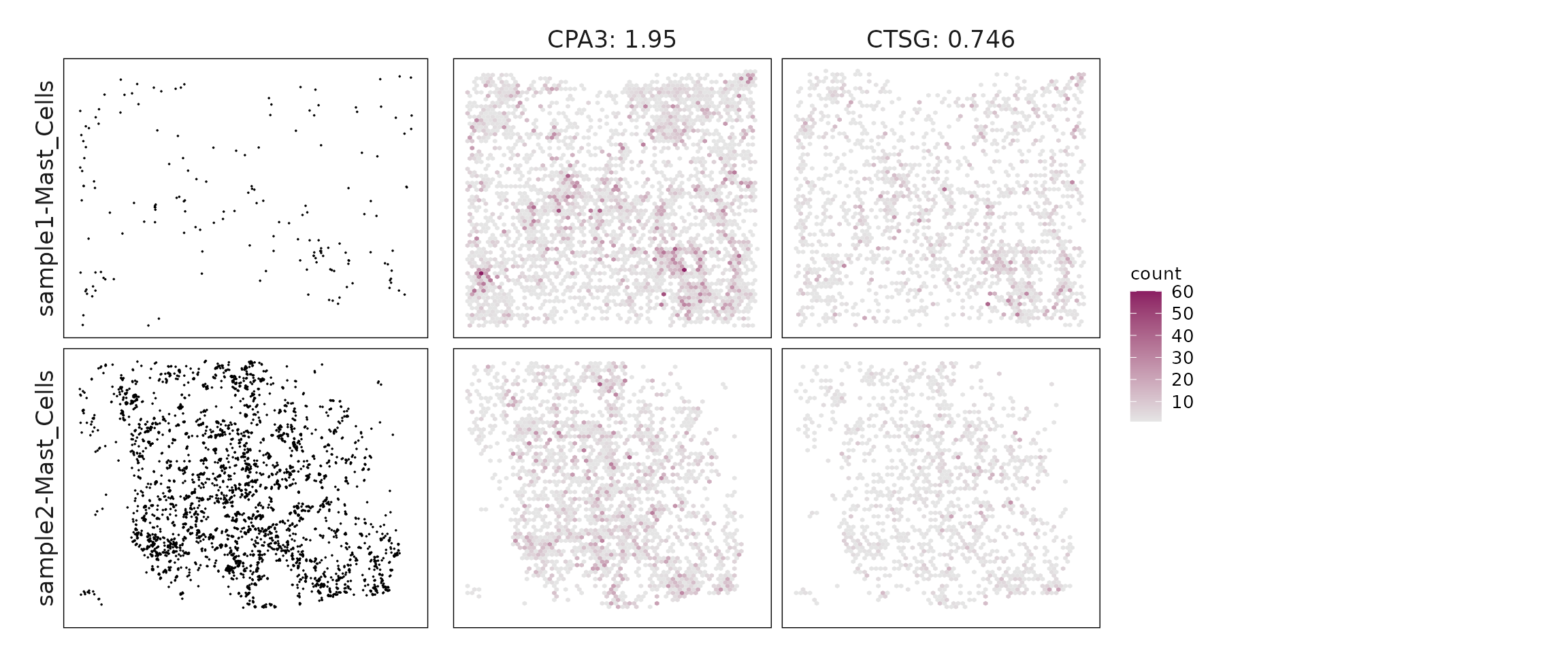
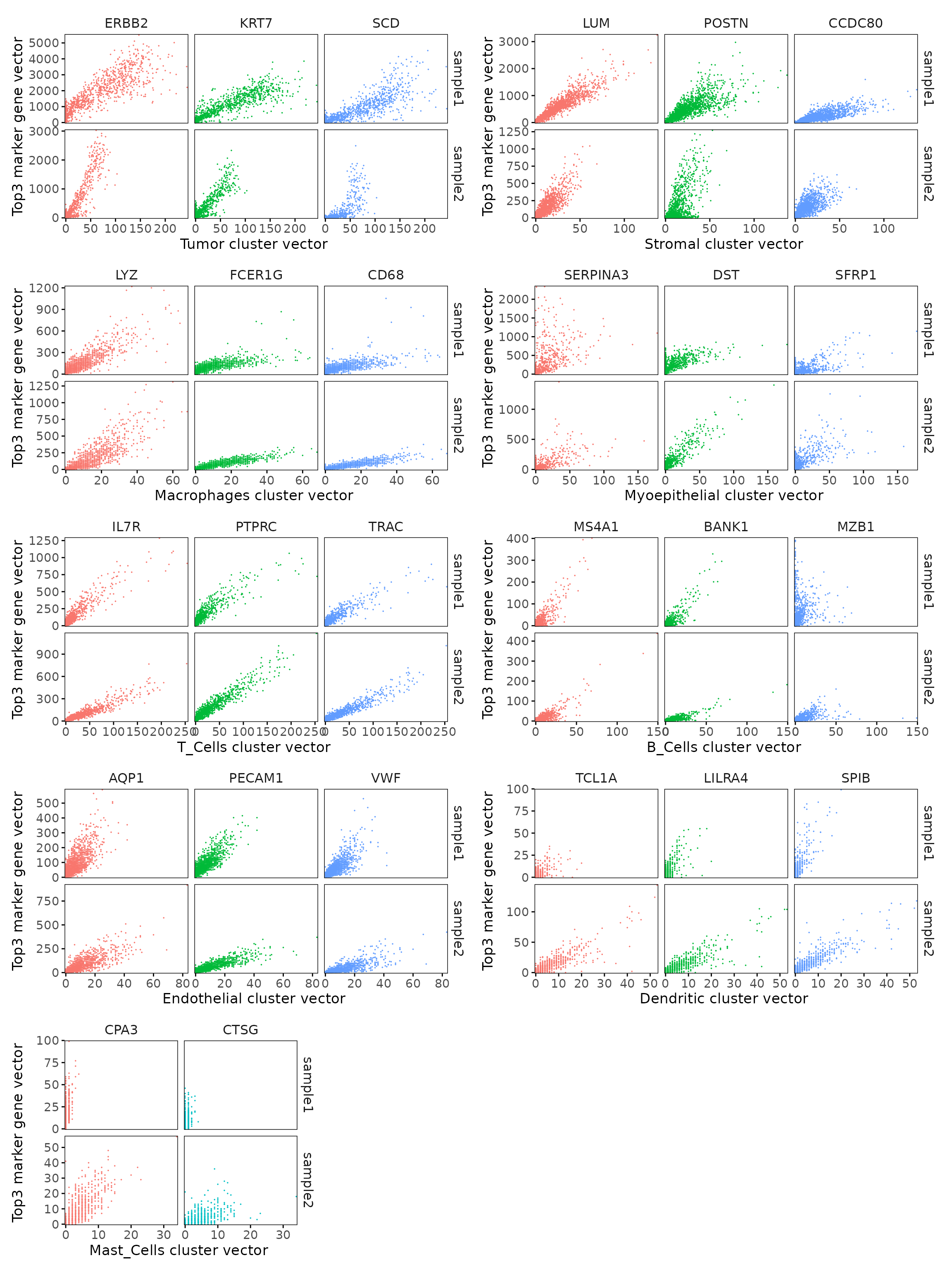
We compared the performance of different marker detection methods — jazzPanda (linear modelling), limma, and Seurat’s Wilcoxon Rank Sum test — by evaluating the cumulative average spatial correlation between ranked marker genes and their corresponding target clusters at the spatial vector level.
plot_lst=list()
cor1 <- cor(sv_lst$gene_mt[1:1600, ],
sv_lst$cluster_mt[1:1600, paste0("c", 1:9)], method = "spearman")
cor2 <- cor(sv_lst$gene_mt[1600:3200, ],
sv_lst$cluster_mt[1600:3200, paste0("c", 1:9)], method = "spearman")
cor_M <-(cor1 + cor2) / 2
Idents(seu)=cluster_info$cluster[match(colnames(seu),
cluster_info$cells)]
for (cl in cluster_names){
ct_nm = anno_df[anno_df$cluster==cl, "anno_name"]
fm_cl=FindMarkers(seu, ident.1 = cl, only.pos = TRUE,
logfc.threshold = 0.1)
fm_cl = fm_cl[fm_cl$p_val_adj<0.05, ]
fm_cl = fm_cl[order(fm_cl$avg_log2FC, decreasing = TRUE),]
to_plot_fm =row.names(fm_cl)
if (length(to_plot_fm)>0){
FM_pt = data.frame("name"=to_plot_fm,"rk"= 1:length(to_plot_fm),
"y"= get_cmr_ma(to_plot_fm,cor_M = cor_M, cl = cl),
"type"="Wilcoxon Rank Sum Test")
}else{
FM_pt <- data.frame(name = character(0), rk = integer(0),
y= numeric(0), type = character(0))
}
limma_cl<-topTable(fit.cont,coef=cl,p.value = 0.05, n=Inf, sort.by = "p")
limma_cl = limma_cl[limma_cl$logFC>0, ]
to_plot_lm = row.names(limma_cl)
if (length(to_plot_lm)>0){
limma_pt = data.frame("name"=to_plot_lm,"rk"= 1:length(to_plot_lm),
"y"= get_cmr_ma(to_plot_lm,cor_M = cor_M, cl = cl),
"type"="limma")
}else{
limma_pt <- data.frame(name = character(0), rk = integer(0),
y= numeric(0), type = character(0))
}
lasso_sig = jazzPanda_res[jazzPanda_res$top_cluster==cl,]
if (nrow(lasso_sig) > 0) {
lasso_sig <- lasso_sig[order(lasso_sig$glm_coef, decreasing = TRUE), ]
lasso_pt = data.frame("name"=lasso_sig$gene,"rk"= 1:nrow(lasso_sig),
"y"= get_cmr_ma(lasso_sig$gene,cor_M = cor_M, cl = cl),
"type"="jazzPanda-glm")
} else {
lasso_pt <- data.frame(name = character(0), rk = integer(0),
y= numeric(0), type = character(0))
}
data_lst = rbind(limma_pt, FM_pt,lasso_pt)
data_lst$type <- factor(
data_lst$type,
levels = c("jazzPanda-glm", "limma", "Wilcoxon Rank Sum Test")
)
data_lst$rk = as.numeric(data_lst$rk)
p <-ggplot(data_lst, aes(x = rk, y = y, color = type)) +
geom_step(size = 0.8) + # type = "s"
scale_color_manual(values = c("jazzPanda-glm" = "red",
"limma" = "black",
"Wilcoxon Rank Sum Test" = "blue")) +
labs(title = ct_nm, x = "Rank of marker genes",
y = "Cumulative average correlation",
color = NULL) +
scale_x_continuous(limits = c(0, 50))+
theme_classic(base_size = 12) +
theme(plot.title = element_text(hjust = 0.5),
axis.line = element_blank(),
panel.border = element_rect(color = "black",
fill = NA, linewidth = 1),
legend.position = "inside",
legend.position.inside = c(0.98, 0.02),
legend.justification = c("right", "bottom"),
legend.background = element_rect(color = "black",
fill = "white", linewidth = 0.5),
legend.box.background = element_rect(color = "black",
linewidth = 0.5)
)
plot_lst[[cl]] <- p
}
combined_plot <- wrap_plots(plot_lst, ncol = 3)
combined_plot 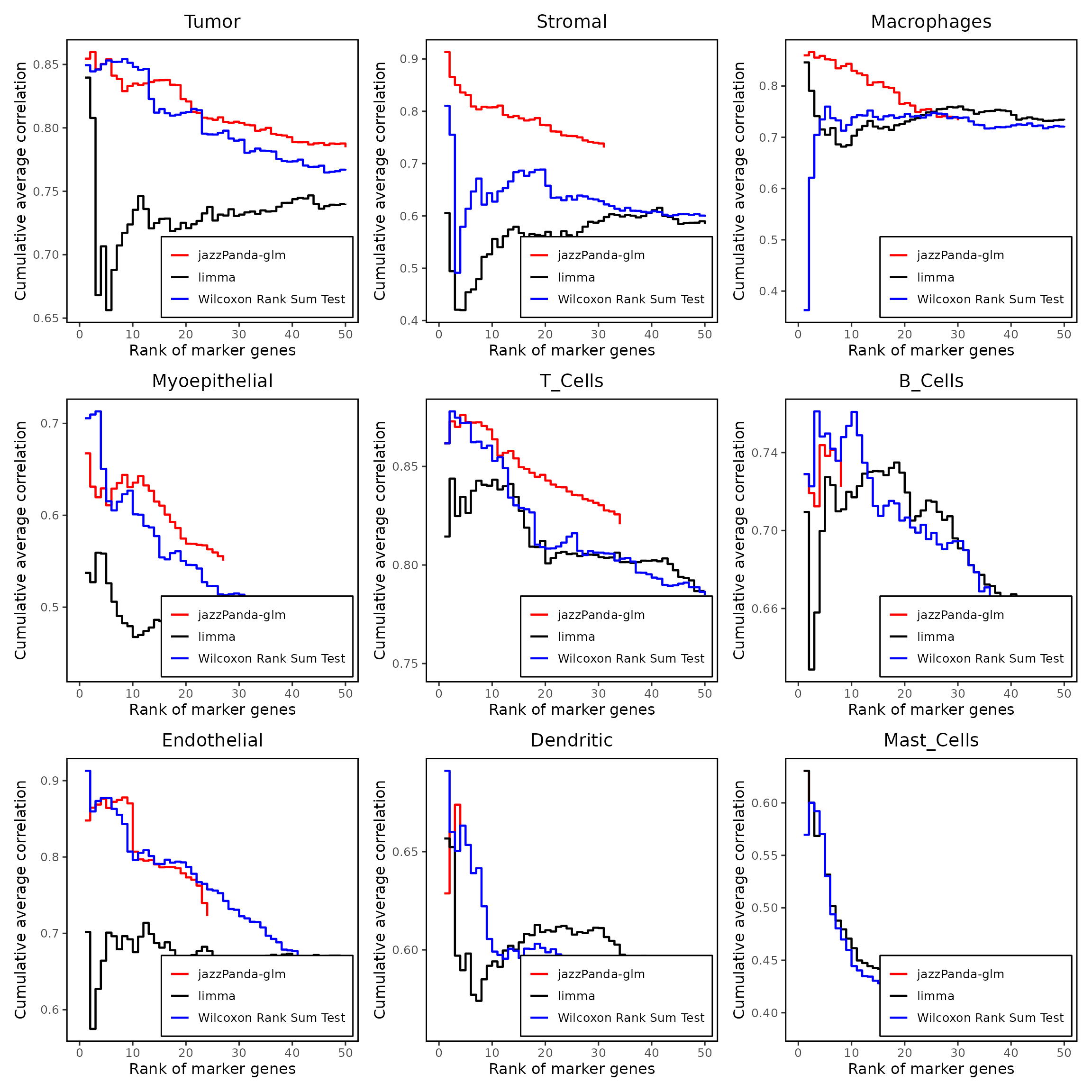
Upset plot
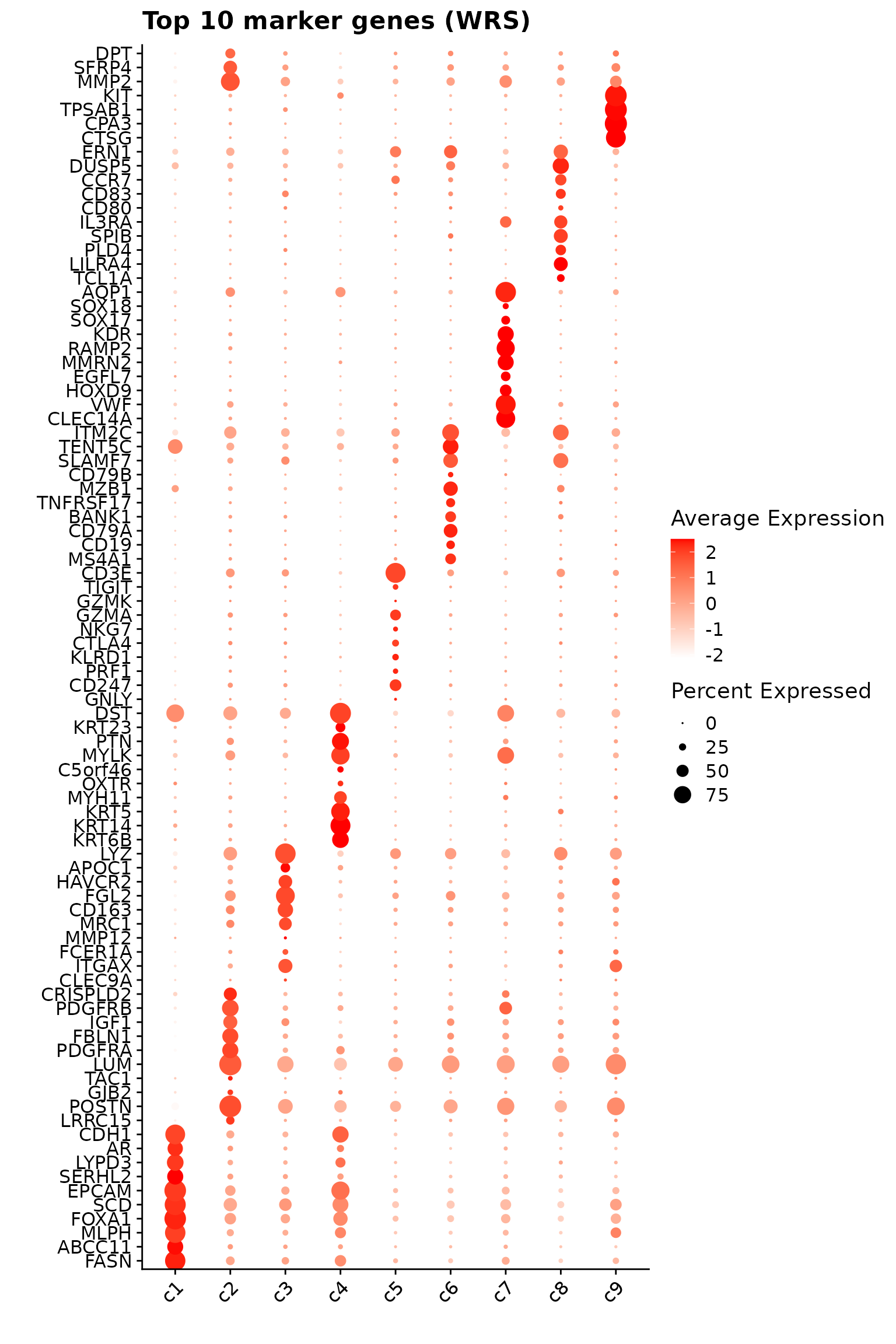
We further visualized the overlap between marker genes detected by the four methods using UpSet plots for each cluster. There is some overlap among methods; however, limma and the Wilcoxon rank sum test identify substantially more marker genes, many of which are not spatially relevant to the clusters. Marker genes for Tumor and T-cell clusters show greater consistency and overlap across methods.
plot_lst=list()
for (cl in cluster_names){
ct_nm = anno_df[anno_df$cluster==cl, "anno_name"]
findM_sig <- FM_result[FM_result$cluster==cl & FM_result$p_val_adj<0.05,"gene"]
limma_sig <- row.names(limma_dt[limma_dt[,cl]==1,])
lasso_cl <- jazzPanda_res[jazzPanda_res$top_cluster==cl, "gene"]
df_mt <- as.data.frame(matrix(FALSE,nrow=nrow(jazzPanda_res),ncol=3))
row.names(df_mt) <- jazzPanda_res$gene
colnames(df_mt) <- c("jazzPanda-glm",
"Wilcoxon Rank Sum Test","limma")
df_mt[findM_sig,"Wilcoxon Rank Sum Test"] <- TRUE
df_mt[limma_sig,"limma"] <- TRUE
df_mt[lasso_cl,"jazzPanda-glm"] <- TRUE
df_mt$gene_name <- row.names(df_mt)
p = upset(df_mt,
intersect=c("Wilcoxon Rank Sum Test", "limma",
"jazzPanda-glm"),
wrap=TRUE, keep_empty_groups= FALSE, name="",
#themes=theme_grey(),
stripes='white',
sort_intersections_by ="cardinality", sort_sets= FALSE,min_degree=1,
set_sizes =(
upset_set_size()
+ theme(axis.title= element_blank(),
axis.ticks.y = element_blank(),
axis.text.y = element_blank())),
sort_intersections= "descending", warn_when_converting=FALSE,
warn_when_dropping_groups=TRUE,encode_sets=TRUE,
width_ratio=0.3, height_ratio=1/4)+
ggtitle(ct_nm)+
theme(plot.title = element_text( size=15))
plot_lst[[cl]] <- p
}
combined_plot <- wrap_plots(plot_lst, ncol = 3)
combined_plot 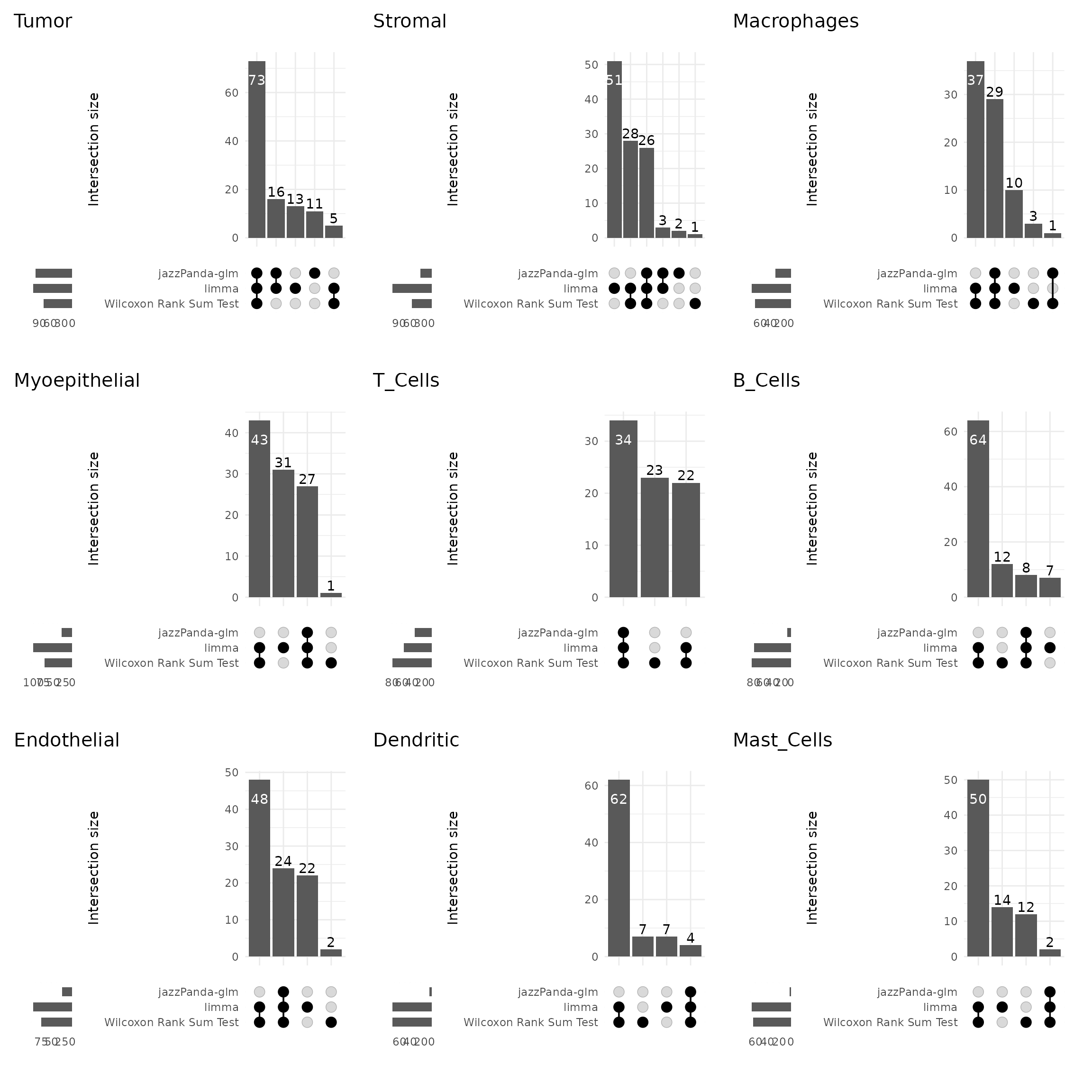
sessionInfo()R version 4.5.1 (2025-06-13)
Platform: x86_64-pc-linux-gnu
Running under: Red Hat Enterprise Linux 9.3 (Plow)
Matrix products: default
BLAS: /stornext/System/data/software/rhel/9/base/tools/R/4.5.1/lib64/R/lib/libRblas.so
LAPACK: /stornext/System/data/software/rhel/9/base/tools/R/4.5.1/lib64/R/lib/libRlapack.so; LAPACK version 3.12.1
locale:
[1] LC_CTYPE=en_AU.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_AU.UTF-8 LC_COLLATE=en_AU.UTF-8
[5] LC_MONETARY=en_AU.UTF-8 LC_MESSAGES=en_AU.UTF-8
[7] LC_PAPER=en_AU.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_AU.UTF-8 LC_IDENTIFICATION=C
time zone: Australia/Melbourne
tzcode source: system (glibc)
attached base packages:
[1] stats4 stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] dotenv_1.0.3 here_1.0.1
[3] speckle_1.8.0 edgeR_4.6.3
[5] limma_3.64.1 RColorBrewer_1.1-3
[7] patchwork_1.3.1 gridExtra_2.3
[9] corrplot_0.95 ComplexUpset_1.3.3
[11] caret_7.0-1 lattice_0.22-7
[13] glmnet_4.1-10 Matrix_1.7-3
[15] dplyr_1.1.4 data.table_1.17.8
[17] ggplot2_3.5.2 Seurat_5.3.0
[19] SeuratObject_5.1.0 sp_2.2-0
[21] SpatialExperiment_1.18.1 SingleCellExperiment_1.30.1
[23] SummarizedExperiment_1.38.1 Biobase_2.68.0
[25] GenomicRanges_1.60.0 GenomeInfoDb_1.44.2
[27] IRanges_2.42.0 S4Vectors_0.46.0
[29] BiocGenerics_0.54.0 generics_0.1.4
[31] MatrixGenerics_1.20.0 matrixStats_1.5.0
[33] jazzPanda_0.2.4 spatstat_3.4-0
[35] spatstat.linnet_3.3-1 spatstat.model_3.4-0
[37] rpart_4.1.24 spatstat.explore_3.5-2
[39] nlme_3.1-168 spatstat.random_3.4-1
[41] spatstat.geom_3.5-0 spatstat.univar_3.1-4
[43] spatstat.data_3.1-8
loaded via a namespace (and not attached):
[1] RcppAnnoy_0.0.22 splines_4.5.1 later_1.4.2
[4] R.oo_1.27.1 tibble_3.3.0 polyclip_1.10-7
[7] hardhat_1.4.2 pROC_1.19.0.1 fastDummies_1.7.5
[10] lifecycle_1.0.4 rprojroot_2.1.0 globals_0.18.0
[13] MASS_7.3-65 magrittr_2.0.3 plotly_4.11.0
[16] sass_0.4.10 rmarkdown_2.29 jquerylib_0.1.4
[19] yaml_2.3.10 httpuv_1.6.16 sctransform_0.4.2
[22] spam_2.11-1 spatstat.sparse_3.1-0 reticulate_1.43.0
[25] cowplot_1.2.0 pbapply_1.7-4 lubridate_1.9.4
[28] abind_1.4-8 Rtsne_0.17 presto_1.0.0
[31] R.utils_2.13.0 purrr_1.1.0 BumpyMatrix_1.16.0
[34] nnet_7.3-20 ipred_0.9-15 lava_1.8.1
[37] GenomeInfoDbData_1.2.14 ggrepel_0.9.6 irlba_2.3.5.1
[40] listenv_0.9.1 spatstat.utils_3.1-5 goftest_1.2-3
[43] RSpectra_0.16-2 fitdistrplus_1.2-4 parallelly_1.45.1
[46] codetools_0.2-20 DelayedArray_0.34.1 tidyselect_1.2.1
[49] shape_1.4.6.1 UCSC.utils_1.4.0 farver_2.1.2
[52] jsonlite_2.0.0 progressr_0.15.1 ggridges_0.5.6
[55] survival_3.8-3 iterators_1.0.14 systemfonts_1.3.1
[58] foreach_1.5.2 tools_4.5.1 ragg_1.5.0
[61] ica_1.0-3 Rcpp_1.1.0 glue_1.8.0
[64] prodlim_2025.04.28 SparseArray_1.8.1 xfun_0.52
[67] mgcv_1.9-3 withr_3.0.2 fastmap_1.2.0
[70] digest_0.6.37 timechange_0.3.0 R6_2.6.1
[73] mime_0.13 textshaping_1.0.1 colorspace_2.1-1
[76] scattermore_1.2 tensor_1.5.1 R.methodsS3_1.8.2
[79] hexbin_1.28.5 tidyr_1.3.1 recipes_1.3.1
[82] class_7.3-23 httr_1.4.7 htmlwidgets_1.6.4
[85] S4Arrays_1.8.1 uwot_0.2.3 ModelMetrics_1.2.2.2
[88] pkgconfig_2.0.3 gtable_0.3.6 timeDate_4041.110
[91] lmtest_0.9-40 XVector_0.48.0 htmltools_0.5.8.1
[94] dotCall64_1.2 scales_1.4.0 png_0.1-8
[97] gower_1.0.2 knitr_1.50 reshape2_1.4.4
[100] rjson_0.2.23 cachem_1.1.0 zoo_1.8-14
[103] stringr_1.5.1 KernSmooth_2.23-26 parallel_4.5.1
[106] miniUI_0.1.2 pillar_1.11.0 grid_4.5.1
[109] vctrs_0.6.5 RANN_2.6.2 promises_1.3.3
[112] xtable_1.8-4 cluster_2.1.8.1 evaluate_1.0.4
[115] magick_2.8.7 locfit_1.5-9.12 cli_3.6.5
[118] compiler_4.5.1 rlang_1.1.6 crayon_1.5.3
[121] future.apply_1.20.0 labeling_0.4.3 plyr_1.8.9
[124] stringi_1.8.7 viridisLite_0.4.2 deldir_2.0-4
[127] BiocParallel_1.42.1 lazyeval_0.2.2 RcppHNSW_0.6.0
[130] bit64_4.6.0-1 future_1.67.0 statmod_1.5.0
[133] shiny_1.11.1 ROCR_1.0-11 igraph_2.1.4
[136] bslib_0.9.0 bit_4.6.0